One of the most powerful applications of flow cytometry is for evaluating drug and vaccine efficacy. Multiparameter flow cytometry relies on the use of fluid systems, fluorescent proteins, and optical systems to detect and collect fluorophore signals. When the technology first became available multiparameter meant 2 colors, today it can mean up to 32 colors. The availability of dyes and antibodies and hardware and software improvements, have made possible the simultaneous analysis of a wide range of parameters. Researchers can probe a single cell for multiple markers, define the composition of a cell population or evaluate protein expression levels. However, the complexity of multiplexed flow cytometry studies creates challenges to generating reproducible and publishable data. Below are design, execution, and analysis best practices to keep in mind when planning flow cytometry experiments.
Sample processing:
One of the advantages of flow cytometry is it can be used to generate a lot of data per cell basis. Researchers can probe a heterogeneous cell population derived from almost any solid tissue or body fluid. Regardless of the source, the cell processing protocol must be optimized to yield a homogenous, single-cell suspension of viable cells. The protocol must therefore be optimized for pH, temperature, and other conditions that will impact cell viability and/or produce autofluorescence. And, it must also yield sufficient numbers of quality cells for analysis. Sample quality directly correlates to data quality. As in every scientific experiment, expert technical skill is the only guarantee for experimental success.
Panel design:
Flow cytometry can be applied to almost any type of study if a fluorescent probe/marker is available for it. However, it requires the correct combination of fluorophores and antibody conjugates titrated to match the expression level of the marker(s) of interest. Titration is a simple but important control for minimizing nonspecific binding and optimizing signal detection to ensure reproducibility. Each panel must include appropriate controls specific to each experiment. Controls are needed to exclude signals from unwanted cell populations and to differentiate signals from dead, damaged/dying cells. While it may be possible to use up to 32 colors, in this case more is necessarily better. With increasing number of fluorochromes, there is some loss of sensitivity due to background and spillover. It is better to design panels with the minimum number of markers needed to address specific research questions. It is also important to optimize the instrument to capture weak signals but exclude background noise. Instrument manufacturers will provide performance test certification. Quality control beyond machine performance is the best practice for producing the highest quality data.
Data acquisition & analysis:
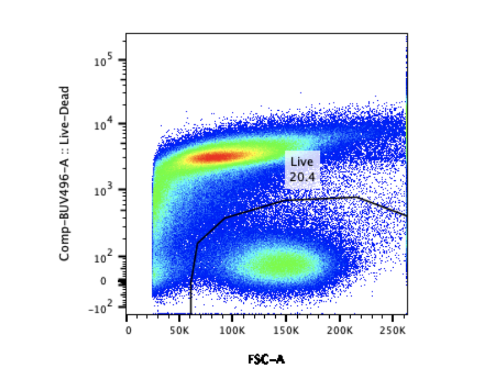
With simultaneous evaluation of several parameters, large volumes of data can be generated from a limited quantity of samples and fast. To be informative the data must capture relevant events and in sufficient numbers. This requires gating strategies that effectively exclude coincidental and background noise. The large and complex data files flow cytometry yields can be both challenging and time-consuming to analyze. Making sense of the seemingly random colored dots requires expert analysis, standardized statistical analysis methods, and access to very expensive statistical software.
Flow cytometry has evolved a great deal since its inception becoming an essential research tool. The design and execution of flow cytometry experiments requires a great deal of technical know-how. Reproducibility is the key to scientific discovery. Continued evaluation and optimization of flow cytometry best practices is the key to generating publishable results.